Detection of Protein Binding sites on DNA Molecules by Solid State Nanopore
Student: Helal Maruf
Degree: Ph.D., December 2022
Major Professor: Dr. Jiali Li
Research Area(s):
Biological Sensors
Biological Materials & Processes
Background/Relevance
-
Several genetic disorders (cancers) and other neurological diseases are found to be associated with protein-DNA binding.
-
Conventional protein-DNA binding detection methods are expensive, DNA read length is limited, imprecise and time consuming.

Innovation
-
Fabricate a new technique using Solid State Nanopore that reduces costs and gets longer DNA read length, label free, tunable pore size.
-
Design high precision, and robust technique.
Approach
- A single nanopore is made to the silicon nitride (SiN) membrane on the platform of a silicon chip. The system is aligned between cis (ground) and trans side of the PDMS connected with KCL electrolyte solution.
- A bias voltage is applied. Biomolecules are passed through the nanopore by the electric field and partially blocks the ion flow, called transient current blockage event, and is then measured and geometry of protein binding sites on DNA molecule is recorded.
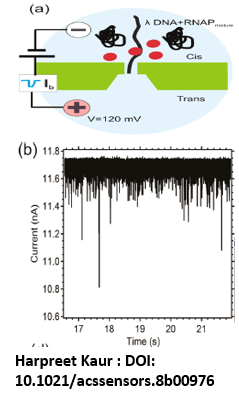
Key Results
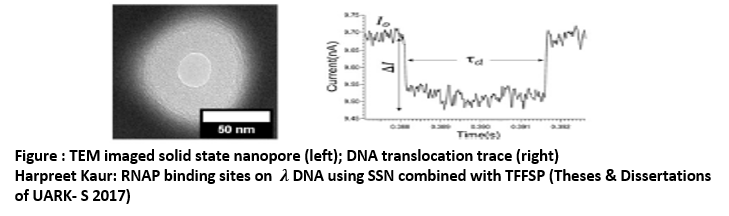
- Fabricated (10-30) nm solid- state nanopore and then imaged by TEM.
- Nanopore device is incorporated with tuning fork force sensing probe tip (TFFSP) system for controlled sensing.
- Translocation of the biomolecules event are determined by the individual binding sites of protein on DNA molecules.
Conclusions
-
The proposed research will quantify amplitude from tip vibration, tip position, and ionic current measurement through solid-state nanopore.
-
Measurement will provide insight of the DNA + Protein_complex from distinct current drop extracted from current blockage data set as a function of frequency, temperature, and applied voltages.
Future Work
-
Future work will be investigating a potential analytical technique in favorable aqueous medium that escalates our comprehension of protein-DNA binding and better cancer diagnosis.
